In my last article, SARS-CoV-2, its components, how it interfaces with the host, and some of the complications it induces upon entering our cells, were explored. In the following sections, I would like to cover a lesser understood proclivity of the virus; its tendency to change and mutate over time, and the challenges associated with the same.
Mentioned previously, SARs-CoV-2 is the name of the virus underpinning sickness/disease (COVID-19) that often, though not always, follows individuals who have become infected. SARS-CoV-2 is an abbreviation for severe acute respiratory syndrome coronavirus 2; “2” was used because the original SARS-CoV occurred in 2003.1 COVID-19 stands for coronavirus disease 2019; 2019 representing the year the disease was identified and characterized.1(1)
STRUCTURE
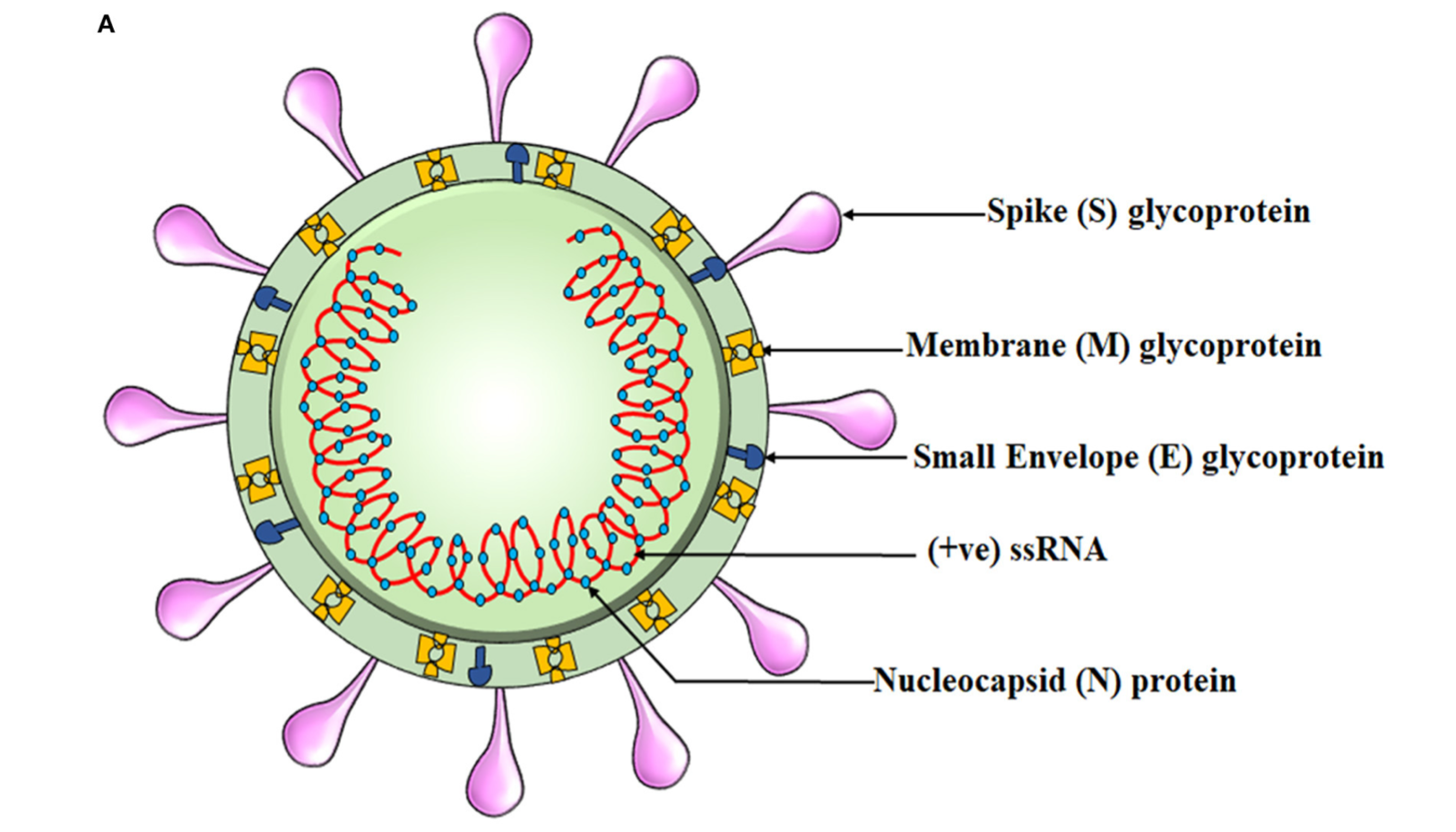
The capsid is essentially a first line of defense against the delicate RNA it holds. The capsid is made from over 400 amino acids; small substances that make larger structures around the RNA, like bricks in a wall.2 See illustration below:
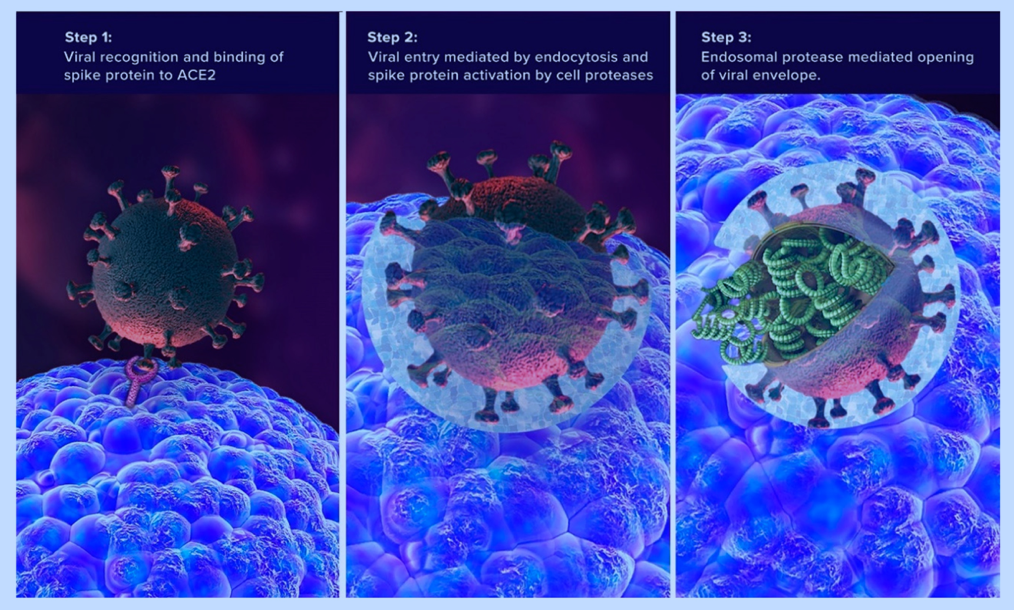
Other components of SARs-CoV-2 include the outer membrane composed of the envelope (E glycoprotein), membrane (M glycoprotein), and the most well-known viral surface protein; the spike protein.3 Similar to the capsid, such proteins are also made of various amino acids and sequences to develop each unique portion of the viral membrane. As a unit, the entire virus with all of its components is termed a virion.4 Of particular interest and relevance is the spike protein; the projections found on the outer surface (see illustration below) each made of 1273 amino acids.5
STRENGTHS OF mRNA VACCINES
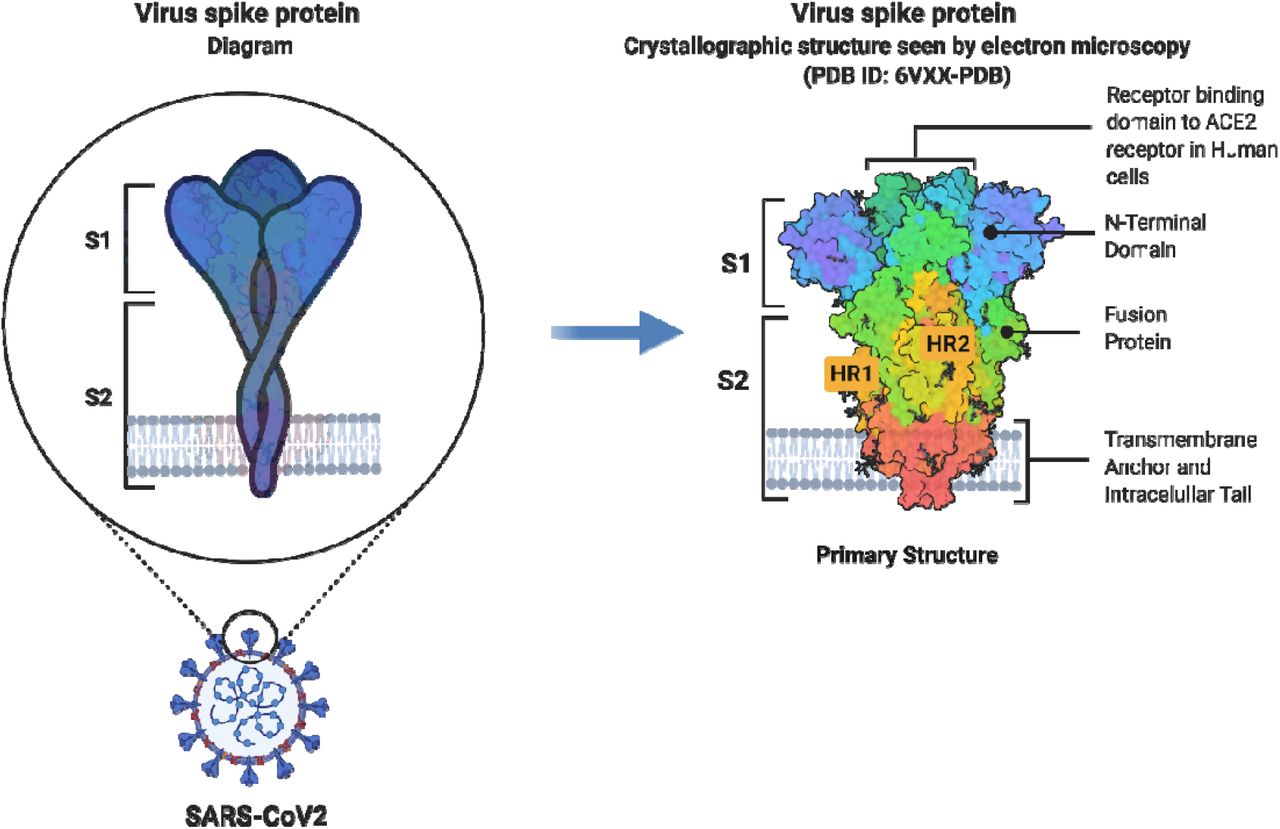
mRNA genetic vaccines are engineered to reproduce the spike protein found on the surface of SARs-CoV-2.6 Spike protein became the point of focus based on evidence from individuals who acquired natural immunity from SARS-CoV-2; they developed antibodies targeted mainly towards the spike protein receptor binding domain (RBD).6(3) Specifically, RBD-specific antibodies were shown to contribute 90% of total neutralizing activity towards said spike protein region in naturally infected individuals.6(3)
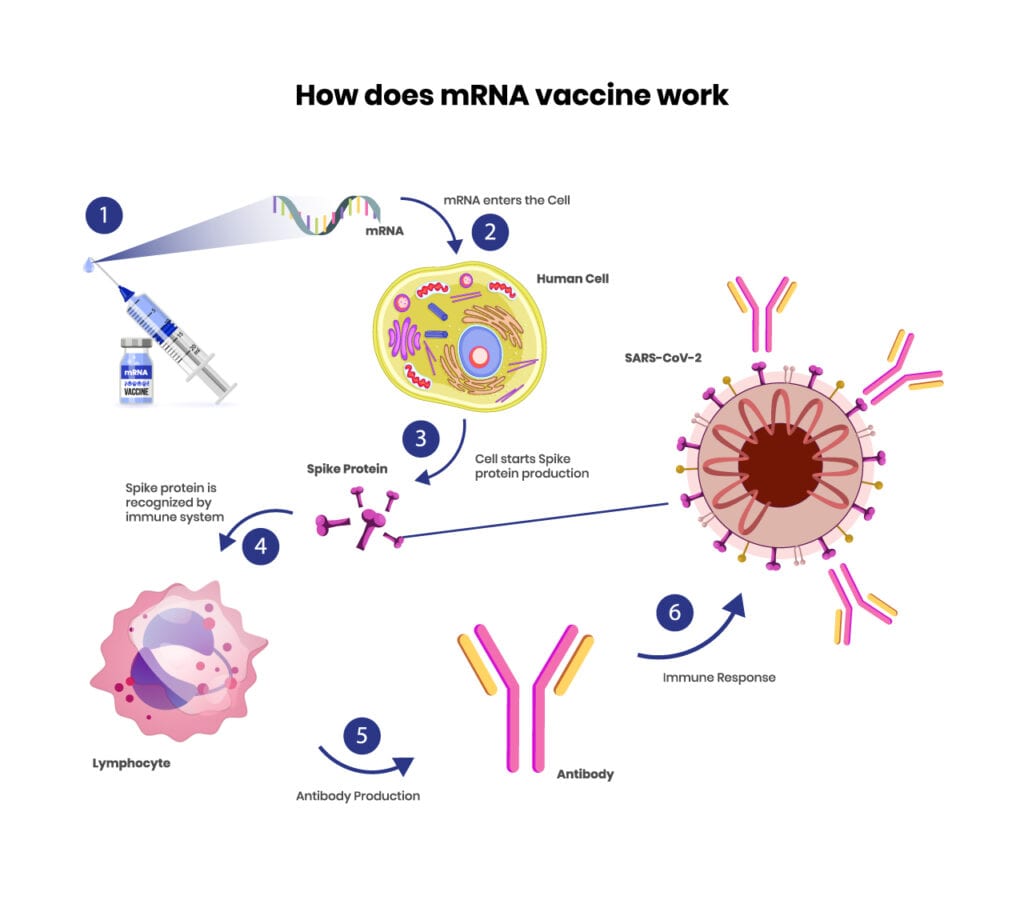
mRNA genetic vaccines are also engineered/intended to anchor themselves on the membrane of the host cell, allowing the innate and adaptive immune system to essentially recognize, interact, and mount a response by synthesizing antibodies (see illustration above).6(3) The process begins by macrophage and dendritic cells (innate immune system) recognizing the presence of the spike protein.7 Essentially, a cascade of events occur thereafter, which ultimately allows for the production of antibodies from B-cells (adaptive immunity).7(e924700-3) Please see the illustration below:
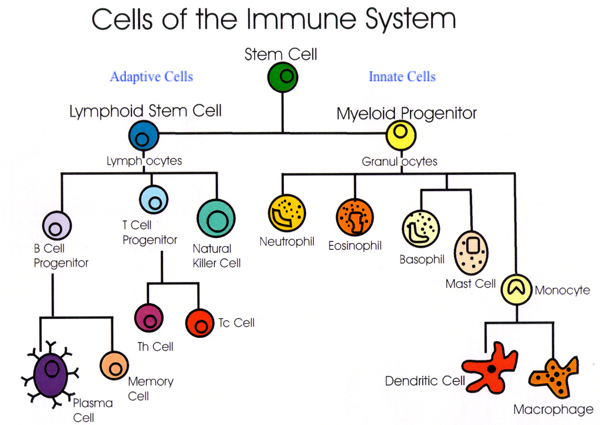
Please see the illustration below for an outline of all main cells of the adaptive and innate immune system:
Antibodies, if they are neutralizing antibodies, will deny entry and hinder the ability of the virus to interact with host cells; in a sense, antibodies might be thought of as “handcuffs” for viruses.7(e924700-3),8 By doing so, neutralizing antibodies also expose the virus to the rest of the immune system; a process that eventually leads to viral destruction and clearance. However, there are some concerns/limitations regarding the mRNA vaccines worth considering.
POTENTIAL LIMITATIONS OF mRNA VACCINES
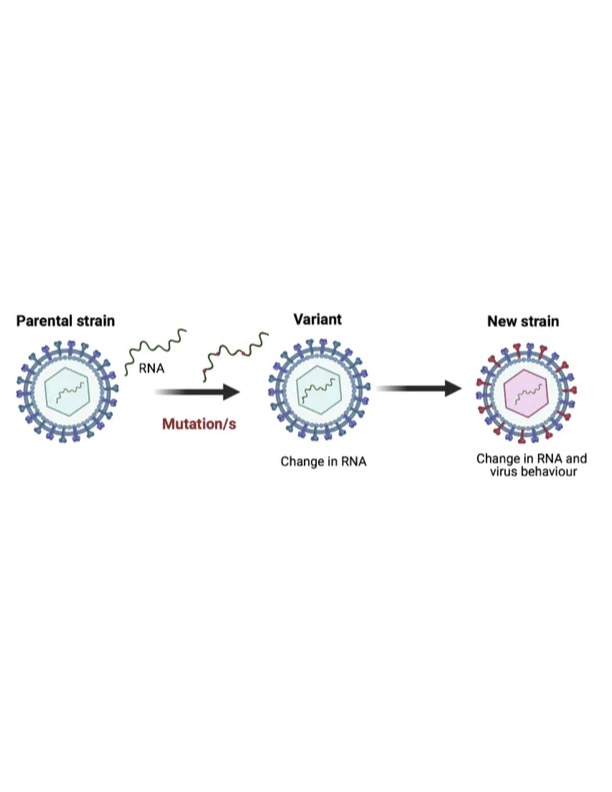
As mentioned in previous sections, mRNA genetic vaccines are engineered to tune the immune system to look for and identify the receptor binding domain (RBD) within the S1 subunit of the spike protein (please refer to the 3rd illustration from the top of the page). SARS-CoV-2 is a single-stranded mRNA virus known to make “offspring” that are not identical to the invading virus (the “parent”) that initiated the replication (copying process making “offspring”).9 Such changes are known as variations, or variants, which can be found amongst a total estimated 109-1011 (or 10,000000000-10,00000000000) virions at peak infection.9(1),10
The offspring become an issue if, during replication, the RBD of the spike S1 subunit is expressed differently enough (different shape, different amino acids) from the original SARS-CoV-2. Such is highly relevant because the mRNA vaccine tunes the individual’s immune system to only recognize the characteristics of the SARS-CoV-2 virus for which it was engineered for.11 Ultimately, variants are camouflaged from the immune system. If the variants escape surveillance of the immune system, they are termed escape variants.12
Harvey et al11(14) stated that there is now unequivocal evidence of the changing antigenicity (ability of immune system to detect and fight a virus) and evolution of the SARS-CoV-2 spike protein. Spike RBDs variations that evade immune surveillance (escape variants) are now at significant frequencies in the global virus population.11(14) Furthermore, there is mounting evidence of variants exhibiting resistance to antibody-mediated immunity elicited by vaccines.11(14)
It might be possible, then, that escape variants will increase both in virulence (severity of disease created) and infectivity (proportion of individuals infected from one individual) from administered mRNA technology training the immune system to target a specific epitope (where antibodies bind to the pathogen) of SARS-CoV-2. In other words, vaccines might/could be inducing a selective pressure towards harsher variants that escape and spread among the population. Please see the below link featuring Dr. Robert Malone; physician, virologist, and inventor of mRNA/DNA vaccine technology covering the same in greater detail:
In conclusion, SARS-CoV-2 is a novel virus that is appearing to become more adaptive, virulent, and infectious through the passage of time. The above sections are by no means an exhaustive outline regarding SARS-CoV-2 and associated mRNA vaccines. However, a basic understanding of the nature, strengths, and limitations of the virus and vaccines arguably remains a worthwhile endeavour, as an informed individual.
References
1. Yuen KS, Ye ZW, Chan CP, et al. SARS-COV-2 and COVID-19: The most important research questions. Cell Bio Sci. 2020;10(40):1-5. doi:https://doi.org/10.1186/s13578-020-00404-4.
2. Surjit M, Lal SK. The Nucleocapsid Protein of the SARS Coronavirus: Structure, Function and Therapeutic Potential. Molecular Biology of the SARS-Coronavirus. 2009;129-151. doi:10.1007/978-3-642-03683-5_9. Accessed October 13, 2021.
3. Mariano G, Farthing RJ, Lale-Farjet SLM, et al. Structural characterization of SARS-CoV-2: Where we are, and where we need to be. Front. Mol. Biosci. 2020. doi: 10.3389/fmolb.2020.605236.
4. Burrell CJ, Howard CR, Murphy FA. Virion Structure and Composition.Fenner and White’s Medical Virology. 2017;27-37. doi:10.1016/B978-0-12-375156-0.00003-5
5. Huang Y, Yang C, Xu XF, et al. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol Sin. 2020;41:1141–1149. doi:https://doi.org/10.1038/s41401-020-0485-4.
6. Heinz FX, Stiasny K. Distinguishing features of current COVID-19 vaccines: Knowns and unknowns of antigen presentation and modes of action. NPJ Vaccines.2020;6(104):1-13. doi: https://doi.org/10.1038/s41541-021-00369-6
7. Wang F, Kream RM, Stefano GB. An evidence based perspective on MRNA-SARS-CoV02 vaccine development. Med Sci Monit. 2020;26: e924700-1–e924700-8. doi:10.12659/MSM.924700.
8. Edara VV, Hudson WH, Xie X, et al. Neutralizing antibodies against SARS-CoV-2 variants after infection and vaccination. JAMA. 2021;325(18):1896-1898. doi:10.1001/jama.2021.4388.
9. Lauring AS, Hodcroft EB. Genetic variants of SARS-CoV-2-What do they mean. 2021;325(6):529-531. doi:10.1001/jama.2020.27124.
10. Sender R, Bar-On YM, Glexier S, et al. The total number and mass of SARS-CoV-2 virions. medRxiv. 2021:1-9. doi:10.1101/2020.11.16.20232009.
11. Harvey WT, Carabelli AM, Jackson B, et al. SARS-CoV-2 variants, spike mutations, and immune escape. Nat Rev Microbiol. 2021;19:409-424. doi:https://doi.org/10.1038/s41579-021-00573-0.
-Michael McIsaac